DNA, Technology, and Florida Strawberries 1
Introduction
Modern breeding programs use DNA technologies to enhance variety improvement. The application of biotechnology takes on many forms, from the detection and tracking of genes associated with an important trait such as yield or disease resistance, to the addition of gene sequences that encode a trait. This latter process is referred to as genetic engineering, or colloquially as "GMO" technology. It is important for consumers, Extension professionals, and teachers to understand how DNA-based technology is applied to Florida strawberries because some of the technologies have never been used and others are central to variety improvement. This article explores the individual technologies and how they are, or are not, applied in strawberry variety development.
Improvement by Conventional Breeding
The UF/IFAS strawberry breeding program has been developing strawberry varieties for nearly 70 years (https://gcrec.ifas.ufl.edu/fruit-crops/strawberries/ufifas-strawberry-varieties/). These varieties are developed using a conventional breeding process of crossing and selection. UF/IFAS varieties are bred to be highly adapted to the weather, soils, and growing practices of central Florida and other winter and early-spring production regions around the world. These varieties (also called "cultivars"), such as 'Florida Radiance' (internationally known as 'Florida Fortuna'), Sweet Sensation® 'Florida127' (registered internationally as FL 09 127), and 'Florida Brilliance' combine high yield and disease resistance with excellent flavor and shelf life (Chandler et al. 2009; Whitaker et al. 2015) (see UF/IFAS Extension publication https://edis.ifas.ufl.edu/hs1322).
Improvement via Genetic Engineering (GMO)
The UF/IFAS strawberry breeding program is often asked, "Are Florida strawberries genetically modified organisms (GMOs)?" The answer is "No." In fact, to date, there has not been a GMO strawberry commercialized anywhere in the world. All commercial strawberry varieties have been developed by conventional breeding methods. While foods derived from genetically engineered crops have shown no evidence of health risks, there are still major social barriers to the acceptance of genetic engineering as applied to fresh fruits and vegetables.
Crops improved with genetic engineering are developed to include a new trait, encoded by a DNA sequence not naturally found in the variety. The gene sequence cannot be created through the conventional breeding process of crossing and selection because it would take decades to add the sequence through breeding alone. Engineered crops often contain DNA sequences from viruses or bacteria that are used to insert the gene of interest or to help it function as desired. The process of insertion of foreign genes into the crop species is called "genetic transformation," and plants developed from this technique are called "transgenic." Transgenic corn, soybean, and cotton varieties are common in the United States (Tester and Langridge 2010). The genetic engineering may add traits that are not naturally present in the species. For example, Bacillus thuringiensis (Bt) corn and cotton are examples of transgenic crops for insect resistance. The Bt gene encoding the information for the bacterial toxin is not naturally present in corn or cotton, so the bacterial gene is inserted via the laboratory (Wallimann 2000; Scriber 2001).
The current crops available in the United States for which some varieties are genetically engineered are canola, cotton, corn, soybeans, alfalfa, sugar beets, Hawaiian papaya, and some varieties of squash. Potatoes and apples have been developed and deregulated, but have limited availability.
Enhancing Variety Development with DNA Markers
In general, the use of conventional breeding approaches to combine many important traits in a single variety is difficult. To make the conventional breeding process more precise and efficient, many crop breeders use DNA technologies to help guide crossing and selection of the best seedlings. Below we describe how the UF/IFAS strawberry breeding program uses these types of technologies to produce better berries.
The UF/IFAS strawberry breeding program has identified certain DNA sequences present at thousands of points along the chromosomes of cultivated strawberry. These DNA sequences can be thought of as the physical addresses of specific chromosome locations, and some will be close by or even inside certain genes of interest. Today, powerful technologies allow the detection of chromosome regions that contain genes controlling a trait. Specific to strawberry, these traits can include disease resistance, fruit quality attributes such as sugar content or aroma, or any other trait that naturally occurs in cultivated strawberry. These gene sequences associated with a trait are valuable because they help breeders follow an important trait without actually measuring the trait. These sequences that are associated with the trait are referred to as "DNA markers" and they allow researchers to follow a trait from generation to generation.
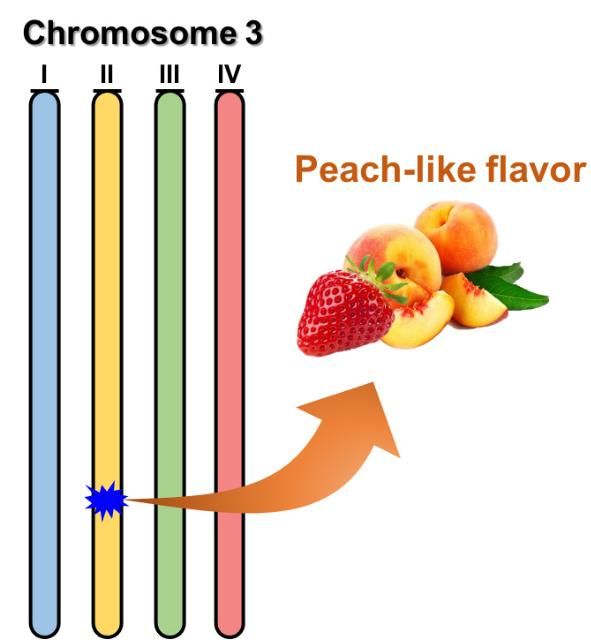
In order to pinpoint the chromosome locations of traits, three things are needed: (1) DNA marker data, (2) observational data on the traits that are carefully measured in the field or lab, and (3) specialized software that can analyze the marker data and the trait data together. At UF/IFAS, we use next-generation DNA sequencing and advanced software called FlexQTL™ that has the ability to trace genes from new seedlings to their parents, grandparents, and beyond through pedigrees. Pedigree-based analysis has already been used to identify several chromosome regions behind resistance to diseases, such as angular leaf spot caused by the bacterium Xanthomonas fragariae (Roach et al. 2016) and resistance to Phytophthora crown rot caused by Phytophthora cactorum (Mangandi et al. 2017). These diseases destroy plants in commercial strawberry production in Florida every year, and genetic resistance is the best way to combat them. An example of a fruit quality trait for which the chromosome regions is known is an aroma compound that gives a "fruity" scent to the strawberry (Chambers et al. 2014). Discovering the chromosome regions behind naturally occurring traits is the first step in using DNA information in conventional strawberry breeding.
High-Throughput Screening of DNA Markers
Using the DNA information gained in the previous step, the UF/IFAS strawberry breeding program can screen large numbers of seedlings soon after they are germinated and select only the seedlings containing the best predicted combination of traits to grow into mature plants and evaluate in the field. Instead of spending lots of time and energy on physically identifying plants that will resist disease and have the desired fruit aroma, the breeder can predict these important traits ahead of time by searching for plants possessing the targeted DNA markers. In 2020, the UF/IFAS breeding program screened over 54,000 seedlings with DNA markers in a three-week period, throwing away 42,000 and keeping 12,000 for further evaluation.
In a practical sense, how do we screen over 54,000 seedlings for multiple traits with DNA markers in just three weeks? A system that could do this must be rapid, accurate, and inexpensive. It is difficult to extract high-quality DNA from strawberry leaves because the leaves contain chemicals that can interrupt the DNA marker screening process, especially when the DNA is quickly and crudely extracted. Recently, the UF/IFAS strawberry breeding program developed a high-throughput DNA extraction and DNA marker testing system (Noh et al. 2016; Oh et al. 2019). This new system can detect tiny genetic variations very quickly and provides an output that is easy to read on a computer monitor. Figure 2 shows a simplified procedure currently used in the UF/IFAS strawberry breeding program. After crossing, the hybrid seeds are germinated to obtain small seedlings. A small leaf disc is punched from the leaf of each seedling, and crude DNA is prepared for DNA marker testing. Only the desired seedlings with the trait of interest are kept and planted in the breeding program nursery, where commercial-type transplants are grown for field evaluation.
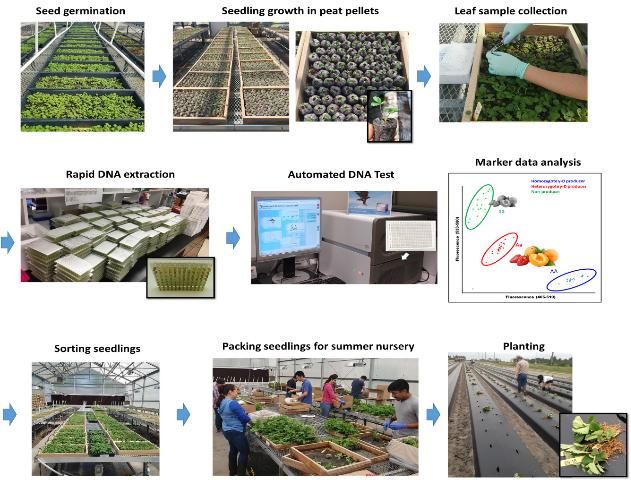
Conclusions
There are many ways to genetically improve plants for the production of new varieties. To date, genetic engineering is not practical and is not used. However, the use of DNA marker technology in strawberry breeding is being used and will continue to increase. At UF/IFAS, more and more strawberry traits will be targeted using the methods described here. These methods are spreading to other strawberry breeding programs in the United States and around the world. We are also working to develop methods to change genes in strawberry by the recent breakthrough technology "gene-editing" (technically known as CRISPR-Cas9), a process that does not pass on the accessory sequences of genetic engineering that some find objectionable. Using this method, changes can be made in discrete genes in elite strawberry varieties quickly, without performing the many generations of crossing that can take decades. These changes are not considered "GMO" by critics because the changes are identical to those that might occur from traditional breeding. As we continue to find more precise and creative ways to breed strawberries, we hope to more rapidly develop strawberry varieties that perform better for the farmer, taste better for the consumer, and have increased disease resistance, making strawberries a healthier crop for Florida farmers and for US consumers.
References
Chambers, A. H., J. Pillet, A. Plotto, J. Bai, V. M. Whitaker, and K. M. Folta. 2014. "Identification of a strawberry flavor gene candidate using an integrated genetic-genomic-analytical chemistry approach." BMC Genomics 15:217. doi:10.1186/1471-2164-15-217
Chandler, C. K., B. M. Santos, N. A. Peres, and C. Jouquand. 2009. " 'Florida Radiance' strawberry." HortScience 44 (6): 1769–1770.
Mangandi, J., S. Verma, L. Osorio, N. A. Peres, E. van de Weg, and V. M. Whitaker. 2017. "Pedigree-based analysis in a multiparental population of octoploid strawberry reveals QTL alleles conferring resistance to Phytophthora cactorum." G3: Genes, Genomes, Genetics 7 (6): 1707–1719.
Noh, Y-H., S. Lee, V. M. Whitaker, K. R. Cearley, and J-S Cha. 2016. "A high-throughput marker-assisted selection system combining rapid DNA extraction and high-resolution melting and simple sequence repeat analysis: Strawberry as a model for fruit crops." Journal of Berry Research 7:1–9. doi:10.3233/JBR-160145
Oh, Y., J. D. Zurn, N. Bassil, P. P. Edger, S. J. Knapp, V. M. Whitaker, S. and Lee. 2019. "The strawberry DNA testing handbook." HortScience 54 (12): 2267–2270.
Roach, J. A., S. Verma, N. A. Peres, A. R. Jamieson, W. E. van de Weg, M. C. Bink , N. V. Bassil, S. Lee, and V. M. Whitaker. 2016. "FaRXf1: A locus conferring resistance to angular leaf spot caused by Xanthomonas fragariae in octoploid strawberry." TAG Theoretical and Applied Genetics 129 (6): 1191–1201. doi:10.1007/s00122-016-2695-1
Scriber, J. M. 2001. "Bt or not Bt: Is that the question?" Proceedings of the National Academy of Sciences of the United States of America 98 (22): 12328–12330. doi:10.1073/pnas.241503398
Tester, M., and P. Langridge. 2010. "Breeding technologies to increase crop production in a changing world." Science 327 (5967): 818–822. doi:10.1126/science.1183700
Wallimann, T. 2000. "Bt toxin: Assessing GM strategies." Science 287 (5450): 41.
Whitaker, V. M., C. K. Chandler, and N. A. Peres. 2015. "Sensation™ 'Florida127' strawberry." HortScience 50 (7): 1088–1091.


