Introduction
Recently, the UF/IFAS strawberry breeding program established somaclonal variation as a method to improve UF strawberries. Somaclonal variation is a breeding method utilizing natural genetic variation induced by a tissue culture process instead of by hybridization. This offers an alternative to mutation breeding for the introduction of new genetic variations in existing strawberry varieties. Preliminary results in the laboratory, the greenhouses, and the field at the UF/IFAS Gulf Coast Research and Education Center (GCREC) suggest that this technique can complement current conventional strawberry breeding methods. Because the spontaneous sequence changes arising in somaclonal variation could be similar to the mutations occurring in nature, crops developed in this way are not regulated as genetically engineered or transgenic plants. The genetic mutations (variations) that occur during the tissue culture process from somatic cells can be used in the development of new cultivars with novel traits. Plant regeneration through this method is relatively fast and easy to screen for the target traits, such as disease resistance and abiotic stress tolerance, without crossings. The main purpose of this publication is to share the potential of this technique with plant breeders in the public and private industries. The secondary purpose is to educate the industry and the public on the scientific background of somaclonal variation.
How Somaclonal Variation Is Different from Other Breeding Technologies
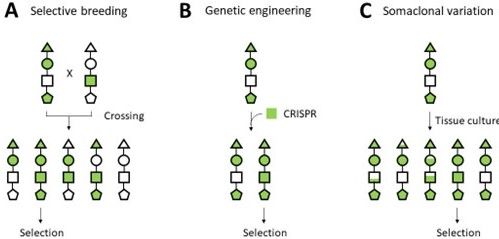
All UF/IFAS strawberry varieties to date have been developed through selective breeding methods. This traditional breeding, or selective breeding, is done by crossing different varieties and selecting the progeny that combines the desirable traits from both parents (Figure 1A). Plant breeders select among the variation resulting from the crosses. In contrast, genetic engineering (Figure 1B) can endow an existing variety with a new trait using specific laboratory techniques. Currently, genetic engineering techniques are not employed in strawberry improvement. Traditional genetic engineering methods introduce or negate the effect of a specific gene. Current gene-editing technologies (CRISPR/Cas9; TALEN) make a few DNA base pair changes, like what could happen in nature, while leaving no foreign DNA behind. Recently, the system of CRISPR gene editing has been established in UF strawberry breeding and testing for new variety development. Additionally, another way to improve a variety is by utilizing somaclonal variation (Figure 1C). Once a new strawberry cultivar is established through selective breeding, the cultivar is propagated by runners (stolons) instead of seeds. This vegetative propagation allows nursery growers to maintain the traits of an elite cultivar through many generations. However, when a small piece of tissue (or explant) (for examples, leaf, petiole, runner, or root) detached from the plant is regenerated through a tissue culture procedure, the process gives rise to noticeable variations in the appearance of the tissue-cultured plant compared to the original variety. This phenomenon is called somaclonal variation, and it is induced by natural mutations that occur during the tissue culture process (Evans 1989). The somaclones (SC) can be screened for desirable breeding traits, such as fruit quality and resistance to biotic (pathogens and insects) or abiotic stresses (heat, drought, and herbicides), to select an improved type. Somaclonal variation, therefore, can bring new traits to the breeding population without crossing.
Credit: UF/IFAS
Somaclonal Variation for Crop Improvement
The genetic variations and diversities are the most important factors of conventional plant breeding programs. Especially, the high level of genetic architecture of traits in strawberry is one of the most essential components to breed superior varieties with good fruit flavor and disease resistance. The typical cycle of developing a new strawberry variety is about 6 years and includes clonal propagations, marker-assisted seedling selection, variety testing and evaluation. Using plant tissue culture-induced somaclonal variation, novel genetic variations can be generated and offered to develop new cultivars. There are advantages and disadvantages to using somaclonal variations for cultivar improvement.
Advantages: This method is cheaper and faster than other genetic transformation methods. It is not necessary to identify genes of the trait cloned for genetic mutation. For example, there must be a known gene function for the use of CRISPR gene-editing tools. Cloning genes in the octoploid strawberry is very difficult without the information of high quality of reference genomes. Additionally, novel DNA sequence variations could be discovered among somaclones, while they are not present in the breeding germplasm. For example, there are no genetic resources available for herbicide tolerance and heat stress resistance in the UF strawberry breeding germplasm. Somaclones raised through cell culture could generate novel traits that are not accessible to conventional breeding.
Disadvantages: One of the major limitations is that somaclonal variations could generate large number of genetic mutations, and it is random and lacks reproducibility. In addition, somaclones have to be validated in field conditions for whether they are heritable and genetically stable. Although a large number of somaclones can be easily generated via in vitro micropropagation, only a limited number of promising somaclones could be selected for the trait of interest. Thus, screening somaclones could be a tedious and labor-intensive process. It has been also reported that the genetic changes occurred from somaclonal variations were not often stable after selfing or crossing (Karp 1992)
Improving Strawberry Varieties by Somaclonal Variation through Tissue Culture
Plant cells have a special ability to develop into a completely new plant under certain conditions. This ability is being utilized in the breeding laboratory to propagate and keep virus-free stock plants. The UF/IFAS breeding program conducts the Strawberry Clean Plant Program (https://edis.ifas.ufl.edu/publication/HS1343) for this purpose. This so-called “meristem culture” is a tissue culture procedure that utilizes meristems, which are the microscopic growing points of the plant. In this case the resulting plant keeps the same genetic traits as the mother plant. This is called direct regeneration.
However, when an explant (meristem or other tissues) is exposed to plant growth hormones (namely auxin and cytokinin), the tissue can generate unorganized masses of cells, or callus. The callus is subsequently reprogrammed to become specific tissue types in the presence of hormones in a culture media (Figure 2). During this developmental reprogramming, combined with stresses from synthetic culture environment, the cells undergo spontaneous DNA mutations. Later, the reprogrammed callus becomes organized to form shoots or induce embryos. Each shoot or embryo becomes a whole new plant (progeny) with subtle variations in its traits that are dictated by the genetic mutations that have occurred. The resulting SCs share almost the same performance as the mother plant, but with some small differences (Zhang et al. 2014). This is called indirect regeneration.
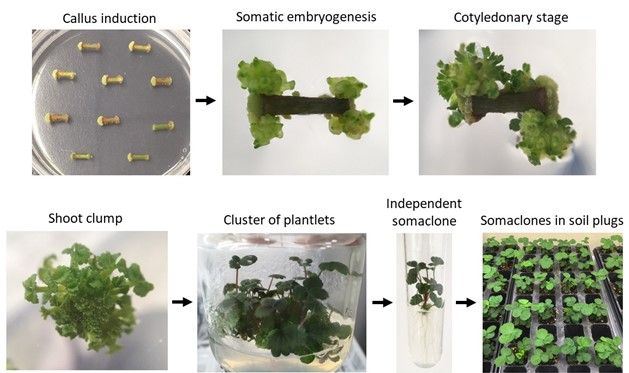

Credit: Cheol-min Yoo, UF/IFAS
Somaclonal variation has been widely used for improvement of traits in diverse crop species (Krishna et al. 2016). To name a few examples, Fusarium wilt (or Panama disease) resistant bananas have been released from somaclonal variation using tissue from the ‘Cavendish’ cultivar, the dominant variety in global production. Somaclonal variation is particularly useful in banana improvement, because banana is difficult to improve via conventional breeding due to the lack of genetic diversity among the commercial varieties (Molina et al. 2016). A second example is carrot leaf blight, which is a common fungal disease of carrot throughout the world. Somaclones from cultivar ‘Fancy’ demonstrated resistance to the disease caused by the fungus Alternaria dauci (Dugdale et al. 2000). Another example is ‘White Baron’ potato released from somaclonal variation of ‘Irish Cobbler’ to retain its white flesh without turning brown after peeling. This trait allowed the industry to expand the prepeeled potato market (Arihara et al.1995).
The UF/IFAS strawberry breeding program has established indirect regeneration protocols for UF varieties of Florida127, Florida Radiance, and Florida Beauty, and will continue to generate protocols for new commercial varieties. In the 2018–2019 season, 250 ‘Florida127’ somaclones were evaluated in the field (Figure 3). Four copies of each clone were tested. The population showed noticeable variation for each SC.
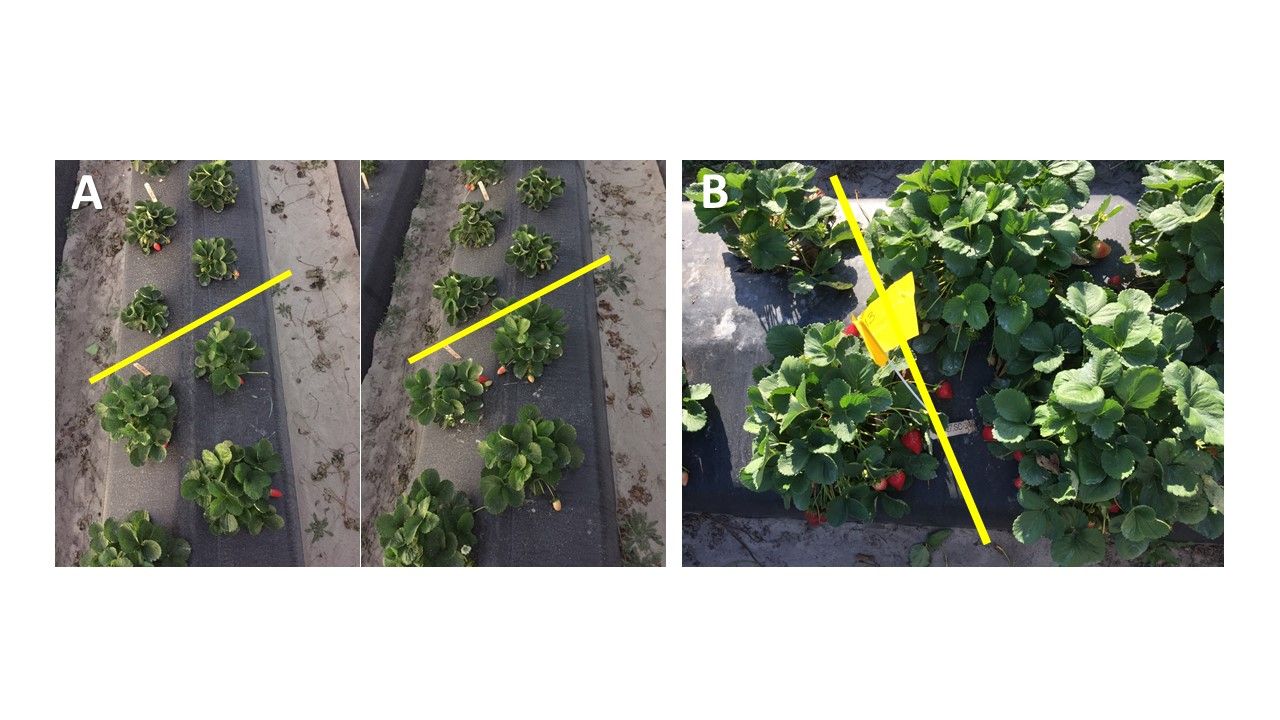

Credit: Seonghee Lee, UF/IFAS
Screening Somaclones for New Breeding Traits
Genetic variability among SCs generated from an indirect regeneration can be exploited as a source for new genetic traits. To isolate a new trait out of a population, it is necessary to be able to screen or select for those that carry the desired trait. One method is to screen a population in the field under normal growth conditions to look for desirable traits like plant height, canopy structure, yield, flowering time, fruit color, flavor, and so on. Another method is to impose a stress on a population and select SCs that are resistant to the stress. The stress can be biotic (bacteria, fungi, parasites, insects etc.), or abiotic (heat, drought, herbicides, etc.). Usually, the latter method allows screening to be done at an earlier developmental stage of the plant. Young plants in a greenhouse in plug trays or even plantlets in test tubes can be screened for the stresses. The degree of stress can be targeted to a mortality rate of 90%–95%, depending on the population size. Then, the resulting 5%–10% survivors become candidates for true resistance to the stress. It is necessary for those candidates to be further screened in field trials to confirm the stress tolerance over generations.
Case Studies in the UF/IFAS Strawberry Breeding
Phytophthora resistance: Phytophthora crown and root rot caused by Phytophthora cactorum is one of the most common and important diseases in Florida strawberry fields. We inoculated 300 and 350 SCs of the susceptible ‘Florida127’ and ‘Florida Radiance’ with spores over multiple seasons. Approximately 7%–8% of the population survived (Figure 4). Some of them are not completely resistant but appear to have increased tolerance. The candidate clones are being tested in the field.
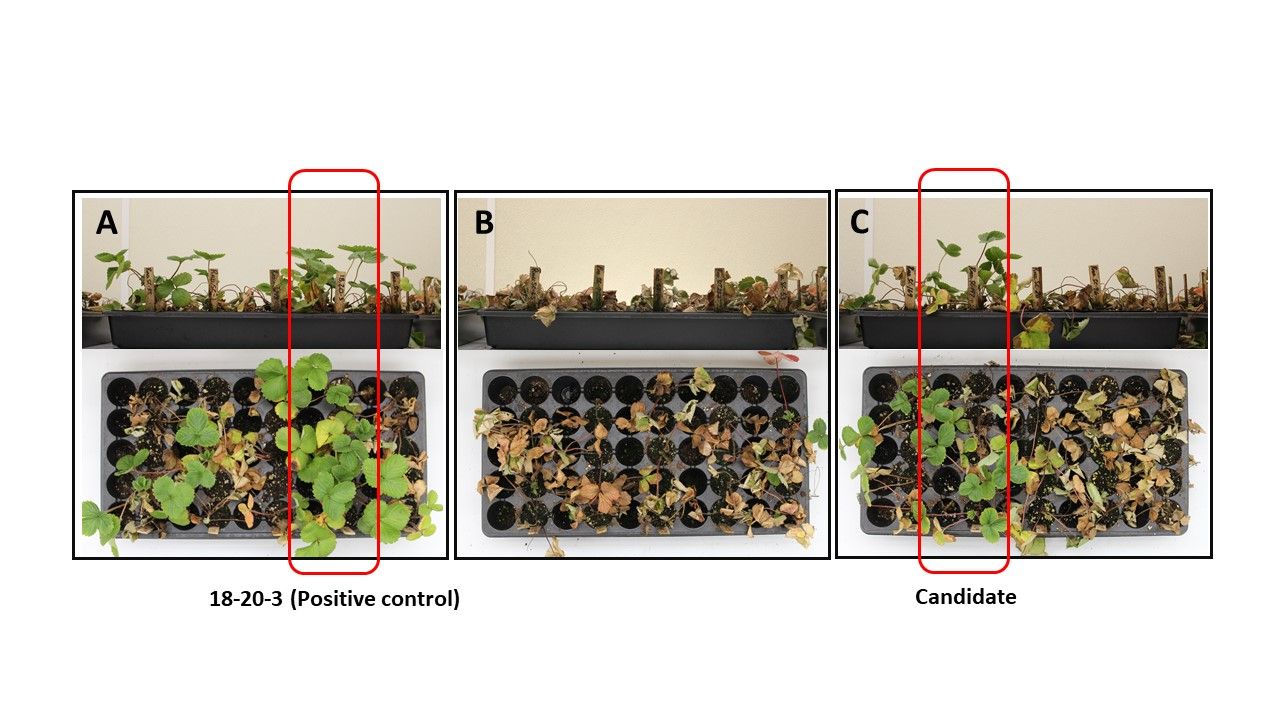

Credit: Cheol-min Yoo, UF/IFAS
Heat stress tolerance: Florida strawberries are planted from late September through mid-October, and high temperatures during this period cause significant challenges. A large amount of irrigation water is needed to cool down the bed temperature and ensure plant establishment. Cultivars that are more tolerant to heat stress can be planted earlier without compromising productivity. We screened 160 SCs of ‘Florida Radiance’ at their seedling stage in 4-inch pots in the greenhouse. The temperature reached around 122°F for three hours a day for 24 days of screening. At the end, the top 20 SCs were selected as tolerant, while 75% of the population showed susceptible symptoms of leaf drying and death. The top four SCs showed normal growth (Figure 5A).
At their plantlet stage in test tube, more somaclone plants can be screened in less space and labor than for the mature plants in the greenhouse. Thus, we tested 500 SCs of ‘Florida Beauty’ in a growth chamber. The chamber was set to 16 hours of light and 8 hours of dark with the temperature of 112°F and 94°F, respectively, for a week of incubation. At the end, 60 SCs survived, and the rest died (Figure 5B).
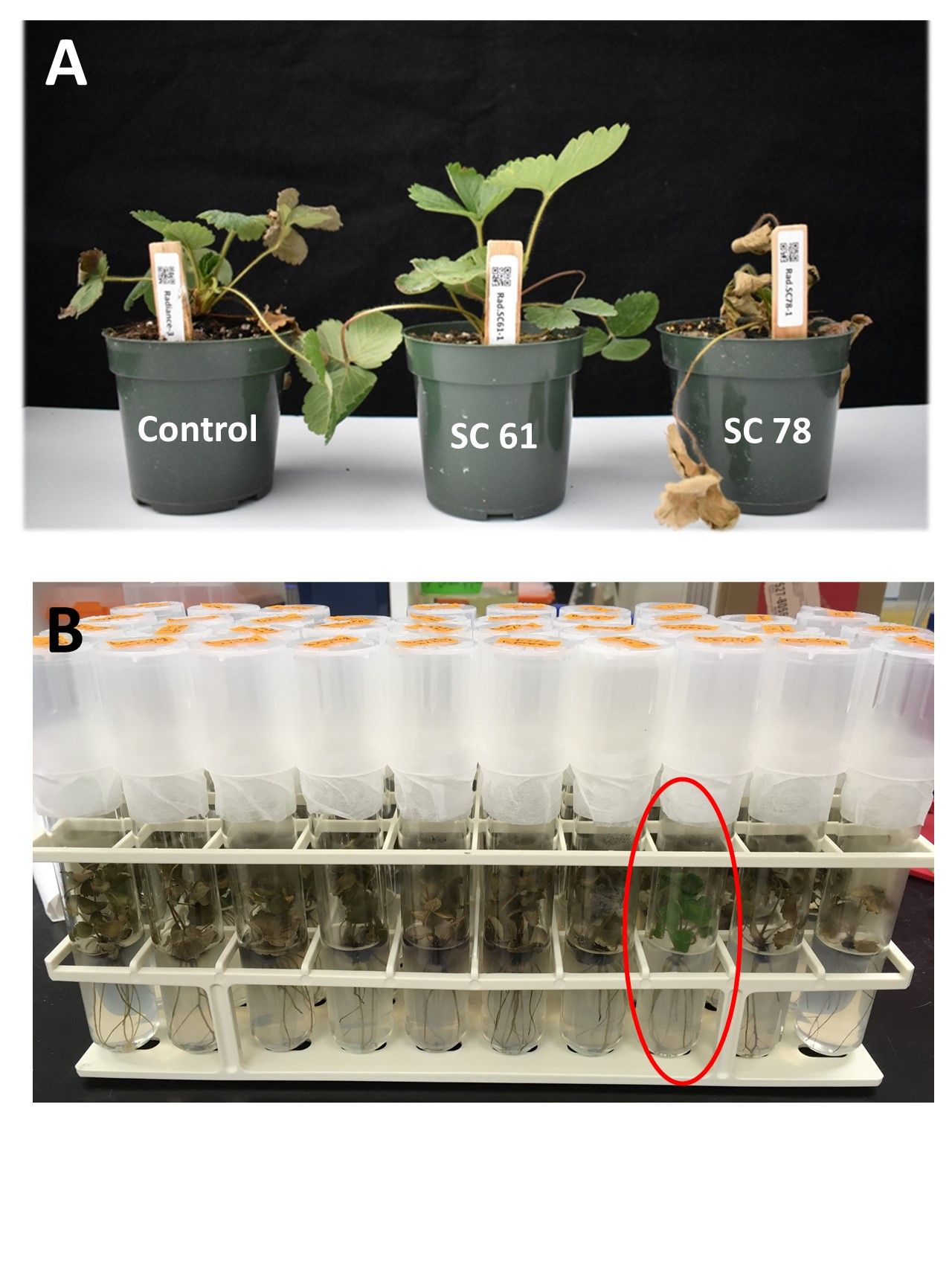

Credit: Cheol-min Yoo, UF/IFAS
Herbicide tolerance: Weed control is important in strawberry fields, requiring herbicide applications. At times, strawberry plants can be damaged by herbicide drift. Here, we tested the possibility of herbicide-tolerant varieties using ‘Florida Beauty’ SCs. A 200-fold diluted solution of Roundup Weather Max® (48.8% glyphosate) was applied to 150 SCs of ‘Florida Beauty’. In this experiment, it was found that a 200-fold dilution is strong enough to kill 99.9% of the somaclone population. The toxicity could be higher when the glyphosate is applied under in vitro conditions rather than application in the field (22 oz/acre). After 3 weeks of incubation, all somaclones were dead except one somaclone that survived without any apparent damage or leaf yellowing (Figure 6).
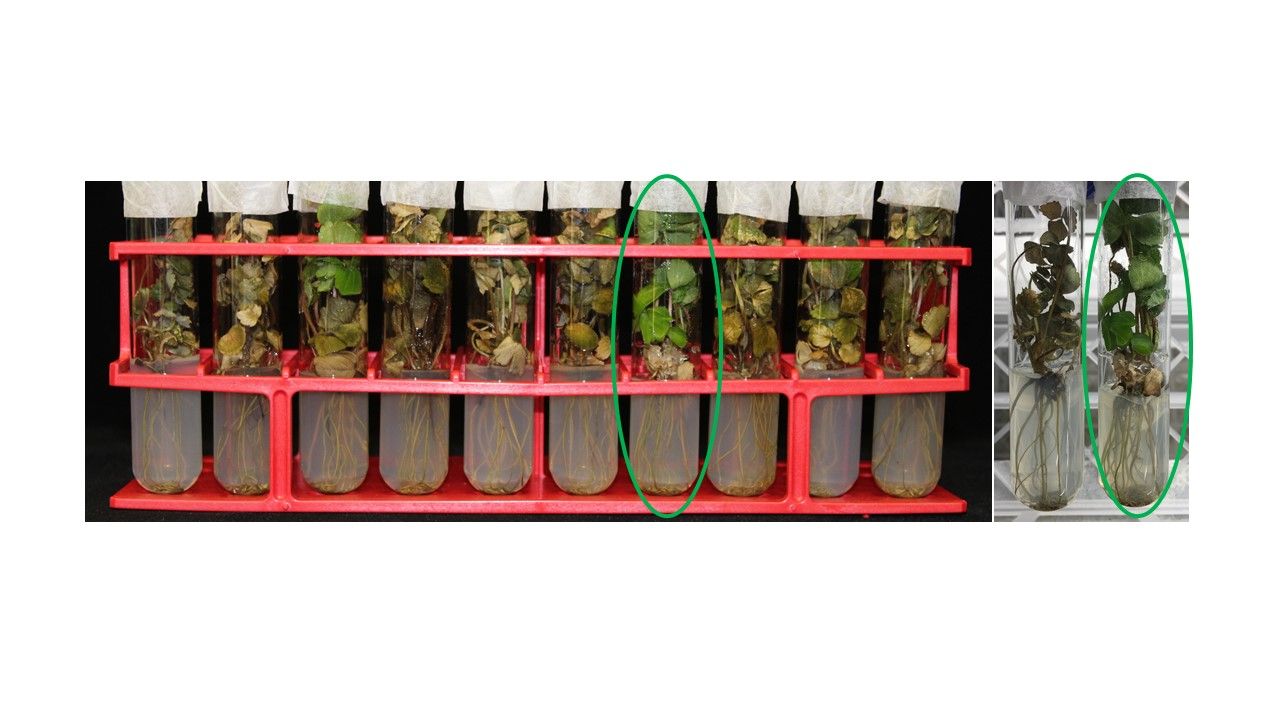

Credit: Cheol-min Yoo, UF/IFAS
Conclusions
Genetic variability induced by somaclonal variation can be effectively used to generate new traits for developing a new strawberry cultivar. This variation is induced by genetic mutation triggered by a tissue culture procedure; thus, varieties produced in this way are not regulated as genetically engineered. A great advantage of this method is that new traits, such as herbicide tolerance and disease resistance to Neopestalotiopsis, can be introduced to the UF strawberry breeding germplasm. However, precise control over the selection pressure and accurate detection methods are required for this technique to be utilized effectively. Additionally, strawberry variety responses to different plant growth regulators would be valuable to determine for the development of somatic variations. It is also important to validate that a new trait is heritable and genetically stable. Because conventional breeding is time-consuming and challenging to improve target breeding characteristics, genetic mutations through the method of somaclonal variations could provide the opportunity to induce DNA sequence changes and introduce target breeding characteristics of interest without altering the genetic combinations.
References
Arihara, A., T. Kita, S. Igarashi, M. Goto, and Y. Irikura. 1995. “White Baron: A Non-Browning Somaclonal Variant of Danshakuimo (Irish Cobbler).” Am Potato J 72 (11): 701–705. https://doi.org/10.1007/BF02849179
Dugdale, L. J., A. M. Mortimer, S. Isaac, and H. A. Collin. 2000. “Disease Response of Carrot Somaclones to Alternaria dauci.” Plant Pathology 49 (1): 57–67. https://doi.org/10.1046/j.1365-3059.2000.00389.x
Evans, D. A. 1989. “Somaclonal Variation—Genetic Basis and Breeding Applications.” Trends Genet. 5 (2): 46–50. https://doi.org/10.1016/0168-9525(89)90021-8
Karp, A. 1992. “The Role of Growth Regulators in Somaclonal Variation.” Br Soc Plant Growth Regul Annu Bull. 2:1–9.
Krishna, H., M. Alizadeh, D. Singh, U. Singh, N. Chauhan, M. Eftekhari, and R. K. Sadh. 2016. “Somaclonal Variations and Their Applications in Horticultural Crops Improvement.” 3 Biotech 6:54. https://doi.org/10.1007/s13205-016-0389-7
Molina, A .B., V. O. Sinohin, E. G. Fabregar, E. B. Ramillete, M. M. Loayan, and C. P. Chao. 2016. “Field Resistance of Cavendish Somaclonal Variants and Local Banana Cultivars to Tropical Race 4 of Fusarium Wilt the Philippines.” Acta Hortic. 1114 (31): 227–230. https://doi.org/10.17660/ActaHortic.2016.1114.31
Zhang, D., Z. Wang, N. Wang, Y. Gao, Y. Liu, Y. Wu, Y. Bai, Z. Zhang, X. Lin, Y. Dong, X. Ou, C. Xu, and B. Liu. 2014. “Tissue Culture-Induced Heritable Genomic Variation in Rice, and Their Phenotypic Implications.” PLoS ONE 9 (5): e96876. https://doi.org/10.1371/journal.pone.0096879